New Publication: Filtering Out the Noise - Metagenomic Classifiers Optimize Ancient DNA Mapping
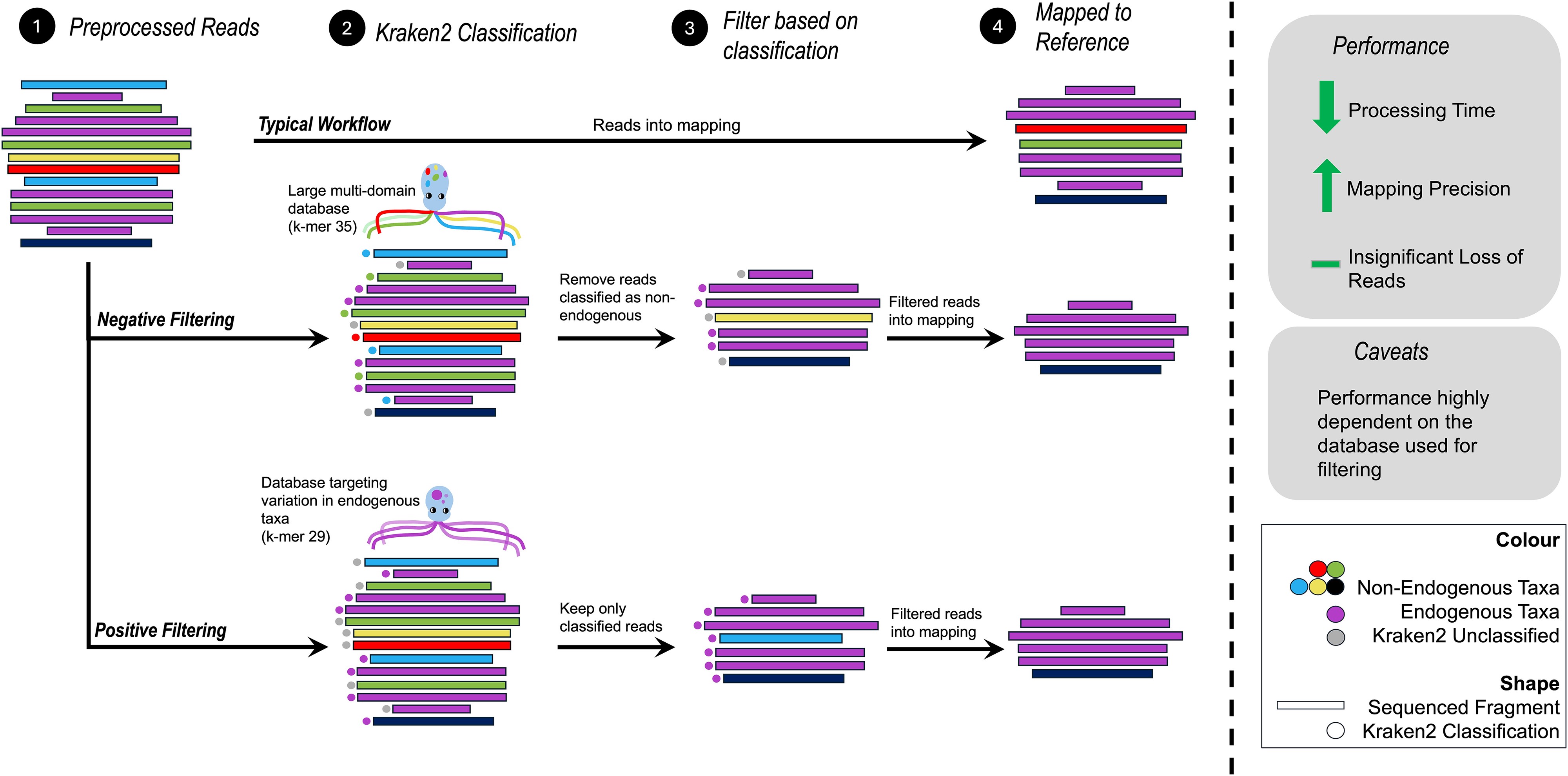
We’re thrilled to announce our new publication in Briefings in Bioinformatics, led by Shyamsundar Ravishankar. This work introduces an innovative workflow that dramatically improves ancient DNA analysis by using metagenomic classifiers to filter contaminating sequences before mapping.
Our approach significantly reduces computational resources (up to 94% reduction in runtime), improves mapping precision, and makes ancient DNA analysis accessible on personal computers—not just high-performance computing clusters.

Abstract
Contamination with exogenous DNA presents a significant challenge in ancient DNA (aDNA) studies of single organisms. Failure to address contamination from microbes, reagents, and present-day sources can impact the interpretation of results. Although field and laboratory protocols exist to limit contamination, there is still a need to accurately distinguish between endogenous and exogenous data computationally. Here, we propose a workflow to reduce exogenous contamination based on a metagenomic classifier. Unlike previous methods that relied exclusively on DNA sequencing reads mapping specificity to a single reference genome to remove contaminating reads, our approach uses Kraken2-based filtering before mapping to the reference genome. Using both simulated and empirical shotgun aDNA data, we show that this workflow presents a simple and efficient method that can be used in a wide range of computational environments—including personal machines. We propose strategies to build specific databases used to profile sequencing data that take into consideration available computational resources and prior knowledge about the target taxa and likely contaminants. Our workflow significantly reduces the overall computational resources required during the mapping process and reduces the total runtime by up to ~94%. The most significant impacts are observed in low endogenous samples. Importantly, contaminants that would map to the reference are filtered out using our strategy, reducing false positive alignments. We also show that our method results in a negligible loss of endogenous data with no measurable impact on downstream population genetics analyses.
Key Achievements
- Dramatic Speed Improvements: Up to 94% reduction in processing time, with 16-fold speed increases for low-endogenous samples
- Accessible Computing: Workflow can run on personal machines, not just HPC clusters, using databases as small as 2.6-2.9 GB
- Improved Precision: Significantly reduces false positive alignments from contaminating sequences
- Minimal Data Loss: Negligible loss of endogenous data (<4%) with no measurable impact on downstream analyses
- Flexible Strategies: Two complementary approaches (positive and negative filtering) suitable for different resource levels and research goals
Innovation
The key innovation is using the metagenomic classifier Kraken2 to filter sequences before mapping to a reference genome, rather than relying solely on post-mapping filters. This pre-filtering approach:
- Removes contaminating sequences that would otherwise spuriously map to the reference
- Drastically reduces the volume of data that needs to be mapped
- Maintains high sensitivity for endogenous (authentic ancient) DNA
- Works particularly well for highly contaminated samples
Practical Impact
This work makes ancient DNA analysis more accessible to researchers worldwide by reducing the computational barrier to entry. Researchers can now process ancient DNA data on standard workstations rather than requiring access to expensive high-performance computing infrastructure.
📄 Read the Full Paper
💻 Access the Workflow
Journal: Briefings in Bioinformatics
Volume 26, Issue 1
Published: December 14, 2024
DOI: 10.1093/bib/bbae646